Blog
How does clustering work in Nexus Copy Number?
When studying genomic data, we are often looking for similarities and differences among a group of samples. We try to see if there are any patterns and whether the samples group together into subsets based on gain and loss profiles. One way to do this is with a clustering function.
In Nexus Copy Number, clustering is done by clicking on the “Cluster” button on the “Results” tab. Clustering uses the chromosomal location, type of copy number event (gain/loss/normal) and size of aberration (large/no aberration) to create sample profiles.
For example, if we had the following three samples w/ aberrations:
Sample S1 has a gain (G), chr1:1-10M
Sample S2 has a gain, chr1:1-5M, normal/no call/no change (N) from chr1:5-10M
Sample S3 has a gain from chr1:5-10M
These will be split into 2 regions – R1=chr1:1-5M and R2=chr1:5-10M
Sample R1 R2
====== === ===
S1 => G G
S2 => G N
S3 => N G
where N stands for no change, D stands for deletion, and G stands for gain, the vector would have five dimensions corresponding to dividing it up as
N NNN D D DD
G NNN N D NN
N NNN N N NN
Each sample profile is converted into a vector. Each dimension of the vector is a region that is constant across all samples. So for example if you have 3 samples, with calls in chr1 having these profiles: NNNNDDDD GNNNNDNN NNNNNNNN
Then it puts a value of 0 in for unchanged regions, 1 in for regions of gain, 4 for high copy gain, -1 for loss, and -4 for multi copy loss.
This would correspond to 3 vectors – S1=(0,0,-1,-1,-1), S2=(1,0,0,-1,0), S3=(0,0,0,0,0) These vectors are clustered as per the settings in File->Options->Analysis Options->Clustering Settings.
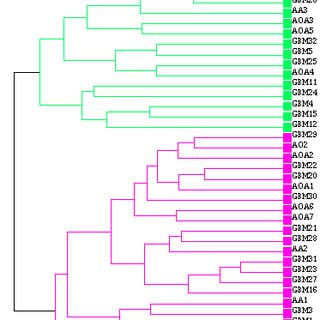
The Hierarchical clustering is bottom up. It takes the two nearest clusters, joins them together, and repeats until there is only a single cluster remaining (the top of the hierarchy). The distance measure is Euclidean (sq. root of sum of squares).
Distances between clusters are computed differently depending on the linkage type selected.
The clustering options in Nexus Copy Number are
Complete Hierarchical (Max): The distance between a pair of clusters A and B is the maximum distance between any two samples a and b where a is in A and b is in B.
Average Hierarchical: The distance between a pair of clusters A and B is the average distance between any two samples a and b where a is in A and b is in B. That is, we compute the distance for all possible pairs drawn from A and B and take the average.
Single Hierarchical (Min): The distance between a pair of clusters A and B is the minimum distance between any two samples a and b where a is in A and b is in B.
K-means: Select number of clusters x and assign x cluster centers. Calculate distance between each sample and cluster center and assign each sample to the closest cluster. Repeat distance calculation and moving the cluster center until there is no more movement.
Hierarchical clustering is exploratory while K-means is used when you know the number of clusters in advance.
Clustering is just one of the many analysis tools found in Nexus Copy Number software. To get an overview of how to use some other tools, view this recent webinar.
